Continuing Education 2023 - Thu, 28 Dec 2023
Last January, the Association of Professional Genealogists @APGgenealogy started requiring that members report at least 12 hours of Continuing Education each year. I found the task of listing my CE time for 2022 quite interesting and last January I posted what I had done.
Below is my Continuing Education activity list for 2023. Each event was 1 hour unless otherwise noted.
Webinars – Total 25.5 hours
- Jan 4 – The 5 steps to organizing your DNA in 2023 – Diahan Southard
- Jan 19 – The Basics of Jewish American Genealogy, Rhonda McClure
- Feb 11 – 10 Tips of Successful Online/Onsite Research in Ukraine, Russia and Belarus – Alina Khuda, Virtual Genealogical Association
- Mar 14 – FamilySearch GEDCOM Technical Q&A – Gordon Clarke
- Mar 14 – RootsTech Recap – Daniel Horowitz, MyHeritage
- Mar 28 – New Developments of MyHeritage DNA by Gal Zrihen, MyHeritage
- Mar 29 – Predicting Unknown Close DNA Relationships Just Got Better! Segcm Tool – Andy Lee, Family History Fanatics
- Mar 29 – The Alex Krakovsky Project – Navigating the Wiki to Locate Town Records, JewishGen
- Apr 11 – First Steps First: Rootstech Recap – Daniel Horowitz
- Apr 21 – DNA Roundtable: Relationship Predictors – Leah Larkin (90 min)
- May 25 – Test. Analyze. Repeat: Long-term DNA Strategies for Success – Diahan Southard.
- Jun 16 – Finding Your Ancestors in Canadian Land Records – Tara Shymanski, Legacy Family Tree Webinars
- Jul 14 – Celebrating 2,000 Webinars! plus 10 tips you can use today – Geoff Rasmussen, Legacy Family Tree Webinars
- Aug 8 – Ten MORE Secrets to Using MyHeritage – Daniel Horowitz, Legacy Family Tree Webinars
- Aug 23 – DNA Painter Basics: Strategies to Enhance Your Genealogical Research – Adina Newman, Virtual Genealogical Association
- Oct 2 – Ask the Experts: Katy Rowe-Schurwanz from FamilyTreeDNA – Diahan Southard (30 min)
- Oct 9 – Ask the Experts: Blaine Bettinger – Diahan Southard (30 min)
- Oct 23 - Ask the Experts: DNA Painter – Diahan Southard (30 min)
- Nov 14 – New Updates on Your MyHeritage Family Tree – Uri Gonen
- Nov 16 – Ask the Wife: A Powerful DNA Strategy – Diahan Southard
- Nov 20 – Ask the Experts: Michelle Leonard – Diahan Southard (30 min)
- Nov 28 – The Good News About Historical Newspapers – Daniel Horowitz
- Nov 30 – Organize Your DNA Matches – Kelli Bergheimer
- Dec 9 – Ten Awesome Things You Can Do on WikiTree – Connie Davis, Virtual Genealogy Association
- Dec 12 – The Lastest Developments in Searching Historical Records on MyHeritage – Maya Geier, MyHeritage
- Dec 15 – Landscape of Dreams: Jewish Genealogy in Canada – Kaye Prince-Hollenberg, Legacy Family Tree Webinars
- Dec 20 – Got Old negatives? Scan Them With Your Phone and These 5 (Mostly) Free Apps! Elizabeth Swanay O’Neal – Family Tree Webinars
Conferences (Online) – Total 14 hours
- Mar 2 to 4 – RootsTech 2023
- Getting Started in Jewish Genealogy – Ellen Kowitt
- What’s New at FamilySearch in 2023 – Craig MIller
- Using DNA to Determine Relationships in 2023 – Beth Taylor
- How third-party DNA tools can help with your family history research – Jonny Perl
- Different Ways to Work with Your family Trees – Uri Gonen
- Tracing Your Jewish Roots in Ukraine – Ellie Vance (30 min)
- Using Maps and Gazetteers to Locate the Hometown – Ellie Vance (30 min)
- What’s New in RootsMagic 9 – RootsMagic
- Nov 2 to 5 – WikiTree Symposium and WikiTree Day
- Mastering the Updated Library and Archives Canada Website – Kathryn Lake Hogan
- DNA Consultations at AmericanAncestors.org – Melanie McComb
- DNA Group Projects and WikiTree – Mags Gaulden
- Tech Troubleshooting – What Would You Do? – Thomas MacEntee
- Keep Your Family’s History Safe for the Future – Marian Burk Wood
- Reverse Phasing – What and Why? – Kevin Borland
- Artificial Intelligence (AI) & Genealogy Panel Discussion, Drew Smith, Dana Leeds, Steve Little, Thomas MacEntee, Rob Warthen, Willie
In total, my time for 2023 was 39.5 hours, which is very similar to my 2022 total of 38 hours.
Planning for 2024
Now is a good time to plan in advance your 2024 activities. I like to add them to my calendar as soon as I find something of interest to me that might contain new or updated information.
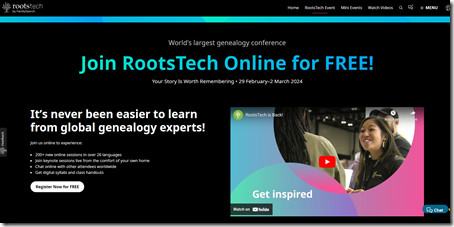
I plan again to attend RootsTech online from Feb 29 to Mar 2.
You can go to their Search the On-Demand Library page and look through their catalog of 4,423 results for over 1,500 sessions from 2019 to 2023 that are still available online. I’m sure you’ll find something of interest there.
They have more than 200 new online sessions planned for 2024. The new session are not yet listed on their site, but they will be soon. When they are ready, you’ll be able to filter your search by year, and 2024 will be an option. Then you can plan the sessions that you’ll want to watch.
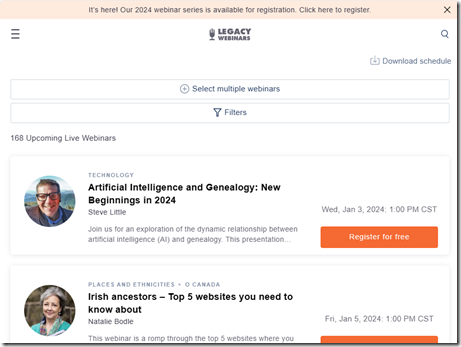
Another planning activity to do right now is to check out which Legacy Family Tree Webinars you’ll want to see in 2024. They just came out with their planned classes and they will feature 112 speakers who will be giving 168 talks. That’s almost one every second day. You can find their list and register for the sessions you want here: Upcoming Webinars - Legacy Family Tree Webinars
I found 17 sessions that I’m already interested in that I’ve now registered for.
Most of the Legacy Family Tree Webinars are free to watch live. Usually, if you miss the live session, you can still watch it for free for about a week.
Of course, another way to get some Continuing Education is to attend a genealogy conference. I have not attended an in-person conference since before Covid. I was planning to finally go on a Genealogy European River Cruise in October 2024 which was to have featured Judy Russell and Blaine Bettinger as the speakers. I was really looking forward to this, but unfortunately it had to be cancelled. It seems I’ll have to wait a while longer until I find another in-person genealogy conference of interest to me.
Now its up to you to get to it. There’s no time like the present to plan some of your genealogical activities for 2024.



 Feedspot 100 Best Genealogy Blogs
Feedspot 100 Best Genealogy Blogs




